Cryo-EM Core
Cryo-EM Core and the David Van Andel Advanced Cryo-Electron Microscopy Suite
 The David Van Andel Advanced Cryo-Electron Microscopy Suite harnesses revolutionary technology to visualize some of life’s smallest — yet most vital — components, including cells and molecular complexes, such as proteins. The Suite encompasses state-of-the-art cryo-electron microscopes (cryo-EM), which are supported by expert staff and a robust high-performance computing cluster with extensive cloud capabilities.
The David Van Andel Advanced Cryo-Electron Microscopy Suite harnesses revolutionary technology to visualize some of life’s smallest — yet most vital — components, including cells and molecular complexes, such as proteins. The Suite encompasses state-of-the-art cryo-electron microscopes (cryo-EM), which are supported by expert staff and a robust high-performance computing cluster with extensive cloud capabilities.
The heart of the Core is the Titan Krios transmission electron microscope, which allows scientists to obtain high-resolution images of cells and molecules in their natural state. The Core also houses a Talos Arctica, often considered the “workhorse” of cryo-EM, and a Tecnai Spirit G2 BioTWIN, which is used for preliminary screening of samples.
The Cryo-EM Core is excited to announced the Cryo-EM Suite is now available for use by external investigators.
For questions about the Suite, please contact Core Manager Dr. Gongpu Zhao via email.
Constructing the Titan Krios
Recent News & Press
Learn More
Cutting thinner, seeing deeper: The power of cryo-PFIB
Tuberculosis is an ancient problem. Modern technology is helping solve it
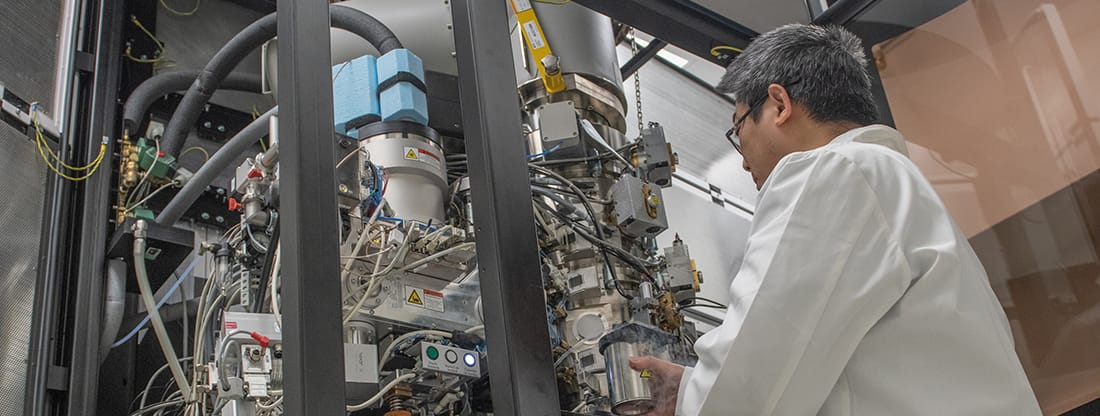

Five years of fueling discovery with cryo-EM
A Ray of Molecular Beauty from Cryo-EM
NIH Director’s BlogVan Andel Institute installs world class microscope to help fight cancer
FOX 17VARI invests $10M in microscope facility
Grand Rapids Business JournalVan Andel Research Institute installs world-class microscopes to enable discovery of the molecular basis of disease
VAIOur Impact
We’re raising thousands to save millions.
We’re turning hope into action for the millions of people around the world affected by diseases like cancer and Parkinson’s. Find out how you can help us make a difference.
- 141 peer-reviewed papers published in 2025, 74 of which were in high-impact journals
- 15 VAI-SU2C Epigenetics Dream Team clinical trials launched to date
- 10 clinical trials and related projects supported by VAI through the International Linked Clinical Trials Program
The David Van Andel Advanced Cryo-Electron Microscopy Suite is a state-of-the-art facility that offers cost-effective cryo-EM and biological EM services along with consulting and hands-on training to internal and external investigators.
Microscopes
Primary Use: Automated, high-throughput production of cryo-lamellae from vitrified cells for cryo-electron tomography (cryo-ET)
The Arctis Cryo-PFIB is a specialized cryogenic plasma-focused ion beam system designed to prepare high-quality, electron-transparent lamellae. It enables precise thinning of vitrified biological specimens while preserving native cellular structures, supporting in situ structural biology workflows.
Key Capabilities
- Cryogenic lamella preparation
- High-precision targeting of regions of interest
- Integrated fluorescence light microscopy (iFLM) for correlative workflows
- Automated, reproducible sample preparation
Specifications
- Electron beam acceleration voltage: 0.2–30 kV
- Multiple ion species: Xenon (Xe), Argon (Ar), Oxygen (O)
- Autoloader capacity: up to 12 cryo-EM grids
- Computer-controlled 4-axis cryo-stage with ±90° tilt
- Triple-beam coincidence (photons, electrons, ions) at the sample position
- iFLM system with 100× objective (NA 0.75)
- 4-channel fluorescence filter set: DAPI, FITC, TRITC, Cy5
Primary use: Data collection
The Titan Krios is a revolutionary, state-of-the-art transmission electron microscope that visualizes cells, proteins and other crucial structures through atomic-resolution 3-D microscopy and cellular tomography. Unlike traditional methods, it does not require samples to be crystallized prior to imaging, saving time and resources while also capturing the target in its natural state.
Capabilities
- 3-D cryo-electron microscopy
- 2-D electron crystallography
- Single particle analysis
- Dual-axis cellular tomography
Specifications
- 80–300 kV operating range
- Gatan Quantum 967 LS imaging filter
- Post-GIF Gatan K3 direct detection camera
- Pre-GIF Ceta 16M camera
- Automated loading for up to 12 samples
- Ultra-stable Schottky field emitter gun
- Three-lens condenser system for parallel sample illumination
- Digital system for remote operation
- Volta phase plate, live image processing
Primary use: Cryo-EM grid screening and data collection
Considered to be the workhorse of VAI’s Cryo-EM Core, the Talos Arctica is a transmission electron microscope that provides high-resolution and high-throughput 2-D and 3-D imaging of frozen, hydrated biological components, including cells, organelles and proteins.
Capabilities
- 3-D cryo-electron microscopy
- 2-D electron crystallography
- Single-particle analysis
Specifications
- 200 kV FEG transmission electron microscope
- Automated loading for up to 12 samples
- Falcon 3EC direct electron detector
- Ceta 16M camera
- K2 direct detection camera
Primary use: Biological sample imaging and protein screening
The Tecnai Spirit G2 BioTWIN is a flexible 80–120 kV microscope mainly used for life science biological specimen imaging.
Capabilities
- Life science TEM imaging
- Negative-stain imaging
- High-contrast, high-resolution imaging
Specifications
- 80–120 kV operational range
- Gatan Orius 832 CCD camera
- Auto-Gun and automatic tuning
- Smart tracking position system
Specifications
- CPU: 2 x Intel Xeon Scalable Gold 5220R, 2.2GHz (24-Core, HT, 2666 MT/s) 150W
- Ram: 768GB PC4-23400 2933MHz
- GPU: 5 x NVIDIA GeForce RTX 3090 TURBO 24G
- Disk: One 16TB SSD
Software
- Cryosparc Live
- Automatic EM and LM tissue processor: Leica EM TP
- Primary use: Automated tissue processing
- Capabilities: Preparation of tissue for embedding
- Ultramicrotome: Leica EM UC7 with EM FC7 cryochamber
- Primary use: Preparation of ultrathin sections at room and cryo temperature
- Capabilities: Preparation of high-quality, consistent ultrathin sections with a thickness feed range from 1 nm–15 µm
- Specifications: Cutting speed 0.05–40 mm/s, feed 1nm–15µm
- Automatic contrast system: Leica EM AC20
- Primary use: Staining of ultrathin section
- Capabilities: Automatic double staining of ultrathin sections
- Specifications: Adjustable stain time and reagent consumption, capable of staining 20 grids at the same time, up to 10 prestored programs
- Compresstome® Vibratome VF-510-0Z
- Primary use: Fully automated vibrating microtome for cutting fresh or fixed tissues with high precision, ideal for electrophysiology, organotypic cultures, imaging, histology and preparing tissue slices for TEM.
- Capabilities: Produces smooth, chatter-free slices from a wide range of tissues while preserving cell viability using agarose embedding and compression technology.
- Specifications: Slice thickness adjustable from ~1 µm to 999 µm (as low as ~4 µm depending on tissue), vibration frequency 0 – 65 Hz, amplitude 1.2 mm, advance speed 0 – 4 mm/s, compatible with stainless steel, ceramic, or tungsten carbide blades
Primary use: Rapidly cryo-immobilizes aqueous samples to preserve ultrastructure, enabling sample preparation for either freeze substitution or cryo-FIB milling.
Capabilities: Freezes cell pellets, C. elegans and tissues.
Specifications: Operating pressure of 2,100 bar. Freeze tissue slices of 100, 200 and 300 µm thick.
Primary use: Performs freeze substitution and progressive lowering of temperature (PLT) techniques.
Capabilities: Maintains precise low-temperature control for dehydration, infiltration, embedding and polymerization using a variety of resins.
Specifications: Operates from –90 °C to +60 °C with ±0.5 °C accuracy, offers up to 35 programmable steps per run, holds multiple vials, and accommodates various embedding carriers and baskets.
Primary use: Characterization of molecular interactions.
Capabilities: Measures binding kinetics, affinity and specificity across a wide dynamic range; supports applications including protein–protein, protein–DNA/RNA, and protein–small molecule interactions.
Specifications: Six flow cells in a single channel; association rate — proteins: up to 3 × 10⁹ M⁻¹·s⁻¹, low molecular weight molecules: up to 5 × 10⁷ M⁻¹·s⁻¹; dissociation rate: 10⁻⁶ to 1 s⁻¹; affinity range: fM to mM.
Other Instrumentation
- Plasma cleaner: Gatan Solarus Model 950
- Plunge freezers: FEI Vitrobot Mark IV
- Carbon coater: HHV BT150
To meet the greater demand for cryo- and biological electron microscopy, we provide a wide range of cost-effective and efficient EM services to VAI investigators, external academic institutes and commercial labs.
- Free cryo-EM project consultation
- Training in all aspects of cryo-EM, including sample preparation, microscope operation, image processing
- Cryo-grid freezing and screening
- Single particle cryo-EM data collection
- Cryo-electron tomography data collection
Training and access to services
Internal users should log in to CrossLab and schedule service times through the system. New users who need login information should contact Lori Moon. Once an account is opened, new users could submit a training request in iLab.
For all Cryo-EM Microscopy inquiries, please contact Dr. Gongpu Zhao.
- Free consultation at the beginning of any new project
- Optimization of experimental procedures and develop methods to meet investigator’s need
- Processing of biological specimens for TEM
- Negative stain TEM imaging
- Immuno-EM
- Ultramicrotome thin sectioning for TEM
- Semi-thin section glass slide preparation and staining for light microscopy
- Production of high-quality ready-to-image samples and high-quality micrographs
- Data interpretation and image analysis
Biological EM workflow
- Typical biological specimens may include tissue blocks, vibratome tissue slices, cell pellets, a monolayer of cells, cells on transwell membranes, etc.
- We can process up to six different types of samples (six different experiments/treatments) per processing run, for the same cost. Multiple specimens per the same type of sample (duplicates) are allowed.
- We do not accept unfixed specimens. We recommend fixing specimens using a fixative that contains both paraformaldehyde and glutaraldehyde (e.g., 2% PFA and 2.5% GA, in 0.1M NaPO4 buffer, pH 7.2-7.4). If perfusion fixation is not possible, get the sample into a fixative solution as quickly as possible once removed from a live source. You can further dice to the appropriate size in fixative.
- Ideally, tissue dimensions for TEM should be ~1mm3. But 1 mm in only one dimension is also acceptable (e.g., 1 x 1 x 2mm). The size limit is due to penetration capability of fixatives for good TEM ultrastructure.
- For cell pellets, we prefer a minimum of 1 million cells per sample. The typical thickness of vibratome tissue sections should be 10–300 um. For negative stain TEM, 1–5 million particles per 100 uL would be acceptable.
- Core staff will perform the sample preparation, ultramicrotomy sectioning, post-staining, and TEM imaging. Once you receive full training on Tecnai G2 Spirit TEM from us, you can perform TEM imaging by yourself.
- Immuno-EM requires special fixation and techniques. Please reach out to us if you are interested in this service.
Core staff will provide a better estimate based on the investigator’s needs. For biological EM services, please contact Dr. Gongpu Zhao and Dr. Ishara Ratnayake.
FAQs
Cryo-EM is short for cryo-electron microscopy, a technique that allows scientists to see tiny molecules down to 1/10,000th the width of a human hair. Although cryo-EM techniques have been around since the mid-1980s, recent advances in technology have led to a scientific revolution, giving scientists the ability to obtain more detailed images of cells, viruses and molecules in their natural state than ever before. In 2015, cryo-EM was named Nature’s Method of the Year.
- An introduction to electron microscopy
- Biological Electron Microscopy books
-
- Biological Electron Microscopy: Theory, Techniques, and Troubleshooting
Michael J. Dykstra and Laura E. Reuss
- Biological Electron Microscopy: Theory, Techniques, and Troubleshooting
-
- Biological Specimen Preparation for Transmission Electron Microscopy
Audrey M. Glauert and P. R. Lewis
- Biological Specimen Preparation for Transmission Electron Microscopy
-
- Electron Microscopy: Principles and Techniques for Biologists
J. Bozzola and L.D. Russell
- Electron Microscopy: Principles and Techniques for Biologists
Structural biology focuses on the shape of the molecules that make up a living organism. As many in the field say, structure is function, meaning that the architecture of a molecule directly informs its role and how it does its job.
Cryo-EM allows scientists to more precisely see proteins and other important structures without many of the constraints posed by traditional methods.
To put its impact in context, in 2007 fewer than 10 molecular structures at less than 5 angstroms (a very tiny unit of measurement) were reported in the literature. Thanks to improvements in cryo-EM technology, by 2015 that number had shot up to nearly 200 (Egelman EH et al; 2016).
Remember, structure informs function. A better understanding of the architecture of cells and molecules gives scientists important insight that allows for the development of new preventative measures, diagnostics and therapies for an untold number of diseases and health problems.
A classic example is that of the key and the lock. If scientists know what the lock looks like, they can develop a key to fit it. In much the same way, if they know what a virus looks like, they are much better equipped to develop a drug that can combat it.
Acknowledgments and Authorship
All work performed by VAI’s Core Technologies and Services should be acknowledged or considered for co-authorship in scholarly reports, presentations, posters, papers and all other publications. Proper acknowledgment allows us to obtain financial and other support that enables us to provide and maintain high-quality support and services. By acknowledging shared resource facilities and instrumentation in publications, presentations and other research communications, you play a critical role in ensuring their continued availability and future development.
Example acknowledgment:
- Cryo-EM data were collected at the David Van Andel Advanced Cryo-Electron Microscopy Suite at the Van Andel Institute (Grand Rapids, MI). We thank the Van Andel Institute Cryo-EM Core (RRID:SCR_023210), especially [staff name], for their assistance with data collection.
HIGHLIGHTED PEER-REVIEWED PUBLICATIONS
Huang Z, Cui W, Ratnayake I, Gallik KL, Cohen L, Tawil R, Pfeifer GP. 2025. SMCHD1 maintains heterochromatin, genome compartments and epigenome landscape in human myoblasts. Nat Commun 16:6900.
Yuan Z, Georgescu R, Yao NY, Yurieva O, O’Donnell ME, Li H. 2024. Mechanism of PCNA loading by Ctf18-RFC for leading-strand DNA synthesis. Science 385(6708).
Hu J, Park SJ, Walter T, Orozco IJ, O’Dea G, Ye X, Du J#, Lü W#. 2024. Physiological temperature drives TRPM4 ligand recognition and gating. Nature.
He Q, Wang F, O’Donnell ME, Li H. 2024. Cryo-EM reveals a nearly complete PCNA loading process and unique features of the human alternative clamp loader CTF18-RFC. Proc Nat Acad Sci U S A 121(18):e2319727121.
Meng X, Ratnayake I, Escobar Galvis ML, Kotecki J, Ramjan Z, Zhao G. 2024. Best practice: setting up and operating a mid-size cryo-EM facility. Front Mol Biosci 10.
Xiao X, Fay A, Santos Molina P, Kovach A, Glickman MS, Li H. 2024. Structure of the M. tuberculosis DnaK−GrpE complex reveals how key DnaK roles are controlled. Nat Comm 15:660.
Li H*, Burgie S*, Gannam ZTK, Li H#, Vierstra R.D#. (2022). Plant phytochromes are asymmetric dimers with unique signaling potential. Nature.
* Co-first authors
# Co-corresponding authors
Zang F, Georgescu RE, Yao NY, O’Donnell ME*, Li H*. (2022). DNA is loaded through the 911 DNA checkpoint clamp in the opposite direction of the PCNA clamp. Nat Struc Mol Biol.
* Co-corresponding authors
Dykstra H, Fisk C, LaRose C, Waldhart A, Meng X, Zhao G, Wu N. 2021. Mouse long-chain acyl-CoA synthetase 1 is active as a monomer. Arch Biochem Biophys 700:108773.
Du M, Yuan Z, Werneburg GT, Henderson NS, Chauhan H, Kovach A, Zhao G, Johl J, Li H, Thanassi DG. 2021. Processive dynamics of the usher assembly platform during uropathogenic Escherichia coli P pilus biogenesis. Nat Commun 12(1):5207.
Yu H, Haja DK, Schut GJ, Wu CH, Meng X, Zhao G, Li H, Adams MWW. 2020. Structure of the respiratory MBS complex reveals iron-sulfur cluster catalyzed sulfane sulfur reduction in ancient life. Nat Commun 11(1):5953.
Xing C, Zhuang Y, Xu TH, Feng Z, Zhou XE, Chen M, Wang L, Meng X, Xue Y, Wang J, Liu H, McGuire TF, Zhao G, Melcher K, Zhang C, Xu HE, Xie XQ. 2020. Cryo-EM structure of the human cannabinoid receptor CB2-Gi signaling complex. Cell 180(4):645–654.e13.
Zhuang Y, Liu H, Zhou EX, Kumar Verma R, de Waal PW, Jang W, Xu TH, Wang L, Meng X, Zhao G, Kang Y, Melcher K, Fan H, Lambert NA, Xu E, Zhang. 2020. Structure of formylpeptide receptor 2-Gi complex reveals insights into ligand recognition and signaling. Nat Commun 11(1):885.
Xu TH, Liu M, Zhou XE, Liang G, Zhao G, Xu HE, Melcher K, Jones PA. 2020. Structure of nucleosome-bound DNA methyltransferases DNMT3A and DNMT3B. Nature 586(7827):151–155.
Bai L, Kovach A, You Q, Hsu HC, Zhao G, Li H. 2019. Autoinhibition and activation mechanisms of the eukaryotic lipid flippase Drs2p-Cdc50p. Nat Commun 10:4142.
Kim JY, Yeom J, Zhao G, Calcaterra H, Munn J, Zhang P, Kotov N. 2019. Assembly of gold nanoparticles into chiral superstructures driven by circularly polarized light. J Am Chem Soc 141(30):11739–11744.
Yu H, Lupoli TJ, Kovach A, Meng X, Zhao G, Nathan CF, Li H. 2018. ATP hydrolysis-coupled peptide translocation mechanism of Mycobacterium tuberculosis ClpB. Proc Natl Acad Sci U S A 115(41):E9560–E9569.
Hu K, Jastrab JB, Zhang S, Kovach A, Zhao G, Darwin KH, Li H. 2018. Proteasome substrate capture and gate opening by the accessory factor PafE from Mycobacterium tuberculosis. J Biol Chem 293(13):4713–4723.
Du M, Yaun Z, Yu H, Henderson N, Sarowar S, Zhao G, Werneburg GT, Thanassi DG, Li H. 2018. Handover mechanism of the growing pilus by the bacterial outer-membrane usher FimD. Nature.
Kang Y*, Kuybeda O*, de Waal PW*, Mukherjee S, Van Eps N, Dutka P, Zhou XE, Bartesaghi A, Erramilli S, Morizumi T, Gu X, Yin Y, Liu P, Jiang Y, Meng X, Zhao G, Melcher K, Ernst OP, Kossiakoff AA, Subramaniam S, Xu HE. 2018. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature.
Fan C*, Choi W*, Sun W, Du J#, Lü W#. 2018. Structure of the human lipid-gated cation channel TRPC3. eLife:e36852.
Yu H, Wu CH, Schut GJ, Haja DK, Zhao G, Peters JW, Adams MWW*, Li H*. 2018. Structure of an ancient respiratory system. Cell.
*Co-corresponding authors
Bai L, Wang T, Zhao G, Kovach A, Li H. 2018. The atomic structure of a eukaryotic oligosaccharyltransferase complex. Nature.
Noguchi Y, Yuan Z, Bai L, Schneider S, Zhao G, Stillman B, Speck C, Li H. 2017. Cryo-EM structure of Mcm2-7 double hexamer on DNA suggests a lagging-strand DNA extrusion model. Proc Natl Acad Sci U S A.
Yang M, Chan H, Zhao G, Bahng JH, Zhang P, Kral P, Kotov NA. 2016. Self-assembly of nanoparticles into biomimetic capsid-like nanoshells. Nat Chem doi:10.1038/nchem.2641
Liu C, Perilla JP, Ning J, Lu M, Hou G, Ramalho R, Himes BA, Zhao G, Bedwell GJ, Byeon IJ, Ann J, Gronenborn AM, Prevelige PE, Rousso I, Aiken C, Polenova T, Schulten K, Zhang P. 2016. Cyclophilin A stabilizes the HIV-1 capsid through a novel non-canonical binding site. Nat Commun 7:10714.
Zhao G*, Perilla JR*, Yufenyuy EL*, Meng X, Chen B, Ning J, Ahn J, Gronenborn AM, Schulten K, Aiken C, Zhang P. 2013. Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Nature 497:643.
*Featured on Nature cover. This paper was selected as NIGMS Director’s Featured Research Advance in June 2013.
Yeom J, Yeom B, Chan H, Smith KW, Dominguez-Medina S, Bahng JH, Zhao G, Chang W, Chang SJ, Chuvilin A, Melnikau D, Rogach AL, Zhang P, Link S, Král P, Kotov NA 2015. Chiral templating of self-assembling nanostructures by circularly polarized light. Nat Mater 14:66–72.
Meng X*, Zhao G*, Yufenyuy E*, Ke D, Ning J, DeLucia M, Ahn J, Gronenborn AM, Aiken C, Zhang P. 2012. Protease cleavage leads to formation of mature trimer interface in HIV-1 capsid. PLoS Pathog 8(8): e1002886.
*Co-first author
Zhao G, Ke D, Vu T, Ahn J, Shah VB, Yang R, Aiken C, Charlton L, Gronenborn AM, Zhang P. 2011. Rhesus TRIM5α disrupts the HIV-1 capsid at the inter-hexamer interfaces. PLoS Pathog 7(3): e1002009.
Byeon IJL, Meng X, Jung J, Zhao G, Yang R, Shi J, Ahn J, Concel J, Aiken C, Zhang P, Gronenborn AM. 2009. Structural convergence between Cryo-EM and NMR reveals novel intersubunit interactions critical for HIV-1 capsid assembly and function. Cell 139:780.
OTHER PEER-REVIEWED PUBLICATIONS
Ruan Z*, Haley E*, Orozco I*, Sabat M, Myers R, Roth R, Du J#, Lü W#. 2021. Structures of the TRPM5 channel elucidate mechanisms of activation and inhibition. Nat Struct Mol Biol.
Ruan Z*, Osei-Owusu J*, Du J, Qiu Z#, Lü W#. 2020. Structures and pH-sensing mechanism of the proton-activated chloride channel. Nature.
Huang Y, Winkler PA, Sun W, Lü W#, Du J#. 2018. Architecture of the TRPM2 channel and its activation mechanism by ADP-ribose and calcium. Nature.
#Co-last authors
Ning J, Erdemci-Tandogan G, Yufenyuy EL, Wagner J, Himes BA, Zhao G, Aiken C, Zandi R, Zhang P. 2016. In vitro protease cleavage and computer simulations reveal the HIV-1 capsid maturation pathway. Nat Commun 7:13689.
Merg AD, Boatz JC, Mandal A, Zhao G, Mokashi-Punekar S, Liu C, Wang X, Zhang P, van der Wel PCA, Rosi NL. 2016. Peptide-directed assembly of single-helical gold nanoparticle superstructures exhibiting intense chiroptical activity. J Am Chem Soc 138(41):13655.
Cassidy C, Himes BA, Alvarez FJ, Ma J, Zhao G, Perilla JR, Schulten K, Zhang P. 2015. CryoEM and computer simulations reveal a novel kinase conformational switch in bacterial chemotaxis signaling. eLife 10.7554/eLife.08419.
Rezikyan A, Jibben ZJ, Rock BA, Zhao G, Koeck FA, Nemanich RF, Treacy MM. 2015. Speckle suppression by decoherence in fuctuation electron microscopy. Microsc Microanal 21(6)1455–1474.
Park J, Nguyen TD, Silveira GQ, Bahng JH, Srivastava S, Zhao G, Sun K, Zhang P, Sharon C, Glotzer SC, Kotov NA. 2014. Terminal supraparticle assemblies from similarly charged protein molecules and nanoparticles. Nat Commun 5:3593.
Song C, Blaber MG, Zhao G, Zhang P, Fry HC, Schatz GC, Rosi NL. 2013. Tailorable plasmonic circular dichroism properties of helical nanoparticle superstructures. Nano Lett 13(7)3256–3261.
Jun S, Zhao G, Ning J, Zhang P. 2013. Correlative microscopy for 3D structural analysis of dynamic interactions. J Vis Exp (76):e50386.
Yang H, Ji X, Zhao G, Ning J, Zhao Q, Aiken C, Gronenborn AM, Zhang P, Xiong Y. 2012. Structural insight into HIV-1 capsid recognition by rhesus TRIM5α. Proc Natl Acad Sci U S A 109(45):18372–18377.
Wang K, Strunk K, Zhao G, Gray JL, Zhang P. 2012. 3D structure determination of native mammalian cells using cryo-FIB and cryo-electron tomography. J Struct Biol 180(2):318–326.
Jun S, Ke D, Debiec K, Zhao G, Meng X, Ambrose Z, Gibson G, Watkins S, Zhang P. 2011. Direct visualization of HIV-1 infection using correlative live-cell microscopy and cryo-EM tomography. Structure 19(11):1573–1581.
Meng X, Zhao G, Zhang P. 2011. Structure of HIV-1 capsid assemblies by cryo-electron microscopy and iterative helical real-space reconstruction. JoVE 54:3041.
Hwang L, Zhao G, Zhang P, Rosi N. 2011. Size-controlled peptide-directed synthesis of hollow spherical gold nanoparticle superstructures. Small 7: doi: 10.1002/smll.201100477.
Song C, Zhao G, Zhang P, Rosi N. 2010. Expeditious synthesis and assembly of sub-100 nm hollow spherical gold nanoparticle superstructures. J Am Chem Soc 132(40):14033–14035.
Zhao G, Treacy MMJ, Buseck PR. 2010. Fluctuation electron microscopy of medium range order in ion-irradiated zircon. Philosoph Mag 90, 4661–4677.
Zhao G, Rougée A, Treacy MMJ, Buseck PR. 2009. Medium-range order in molecular materials: Fluctuation electron microscopy for detecting fullerenes in disordered carbons. Ultramicroscopy 109(2):177–188.
Treacy MMJ, Kumar D, Rougée A, Zhao G, Buseck PR, McNulty I, Fan L, Rau C, Gibson JM. 2008. Glimpsing order within the disarray. J Phys Conf Ser 126(1).
Treacy MMJ, Kumar D, Rougée A, Zhao G, Buseck PR, McNulty I, Fan L, Rau C, Gibson JM. 2007. Probing medium-range structural correlations by fluctuation microscopy. J Phys Condens Matter 19(45).
Zhao G, Zhang Q, Zhang H, Yang G, Tang J, Zhou O, Qin LC. 2006. Field emission of single Cs doped carbon nanotube. Appl Phys Lett 89(26).
Zhao G, Zhang J, Zhang Q, Zhang H, Tang J, Zhou O, Qin LC. 2006. Fabrication and characterization of single nanotube emitters as point electron sources. Appl Phys Lett 89 (19).
Zhang H, Tang J, Zhang Q, Zhao G, Yang G, Zhang J, Zhou O, Qin LC. 2005. Field emission of electron from single LaB6 nanowires. Adv Mater 18, 87.
Zhang H, Zhang Q, Zhao G, Tang J, Zhou O, Qin LC. 2005. Single-crystalline GdB6 nanowire field emitters. J Am Chem Soc 127(38):13120–13121.

Gongpu Zhao, Ph.D.
Director, Cryo-EM Core

Xing Meng, Ph.D.
Core Scientist, Cryo-EM Core

Ishara Ratnayake, Ph.D.
TEM Core Specialist

